 NA-MIC Project Weeks
NA-MIC Project Weeks
Back to Projects List
Small Animal Quantitative Imaging
Key Investigators
- Curtis Lisle (KnowledgeVis, LLC)
- Andrey Fedorov (BWH)
- Yanling Liu (Frederick National Lab for Cancer Research)
Project Description
For this project, we aim to bring small animal MR datasets in DICOM format and repeat the process developed for the QIICR program to segment a lesion (a Neuroendocrine Tumor in this case), convert the segmentation to a DICOM segmentation using the DCMQI slicer extension, and finally measure the segmentation using the Quantitative Reporting module. Our aim is to develop a set of repeatable analysis steps we can put into place to analyze additional datasets in our lab.
Objective
- Develop a set of processing steps for lesion analysis that are repeatable for other small animal datasets. Access if clinical tools from the QIICR program will apply to small animal MR datasets as well.
Approach and Plan
- Test DCMQI tools on a Small Animal MR dataset. Access if small animal DICOM headers are similar enough to clinical scanners.
- Use Slicer segmentation tools to generate a label map of the lesion.
- Create a DICOM segmentation object form the segmented label using DCMQI
- Investigate DeepInfer models for automatic segmentation of later lesions
Progress and Next Steps
- Segment lesion using Slicer Segmentation Wizard
- Follow excellent QIICR tutorial instructions
- DCMQI conversion to DICOM segmentation object failed during the first attempt. Consulted Andrey.
- One of slices from the Phillips small animal scanner was identified with inconsistent header contents compared to other slices.
- Change made to DCMQI to accommodate this dataset.
- Reprocessed successfully and measured DICOM segmentation object using Quantitative Reporting module
- Build and trained a CNN using Keras. Consulted with Alireza about how to connect with DeepInfer.
- Planning to complete DeepInfer integration of our new model over the coming weeks.
Illustrations
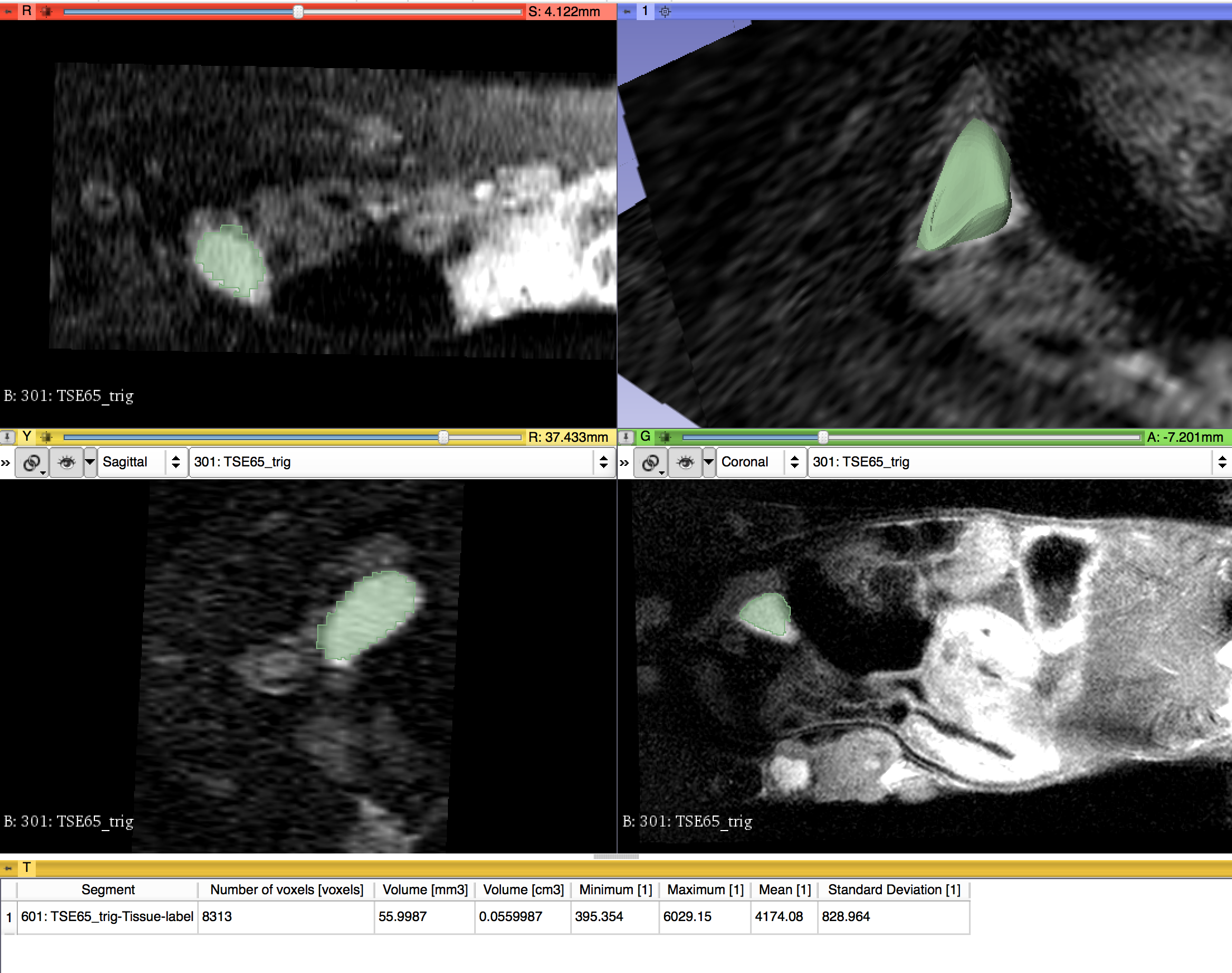
Background and References
- Documentation: http://qiicr.org/dcmqi-guide/tutorials/intro.html
- Documentation: https://qiicr.gitbooks.io/dicom4qi/